Relationship between promoter methylation and protein expression of glutathione S-transferase gene class P1in breast cancer
Supparit Pakdeethai, BSc**, Valla Fongchaiya, BSc**, Tanett Pongtheerat, PhD***,
Kroon-pong Iampenkhae, MD**, Pichet Sampatanukul, MD, MSc**
*This paper was supported by the CU.GRADUATE SCHOOL THESIS GRANT of
Chulalongkorn University.
**Department of Pathology, Faculty of Medicine, Chulalongkorn University, Bangkok, Thailand
*** Unit of Biochemistry, Department of Medical Science, Faculty of Science, Rangsit University,
Patumthani, Thailand
Correspondence : Pichet Sampatanukul, MD, MSc,
Email: fmedpst@gmail.com
ABSTRACT
Background: Glutathione S-transferase gene class P1 (GSTP1) encodes glutathione S-transferase P1 (GSTP1) that catalyses detoxifying metabolism of glutathione. The lack of GSTP1 expression is linked to carcinogenesis and is related to promoter methylation. The previous studies of GSTP1 hypermethylation in breast cancer reveal diverse results and suspicious correlation with GSTP1 expression.
Objective: To investigate the correlation between promoter methylation and protein expression of GSTP1 in breast cancer.
Methods: One hundred breast cancer samples were used. The study of promoter methylation employed meth-ylation specific assay (MSP) on fresh tissues while the assessment of protein expression was done with im-munohistochemistry (IHC) on formalin fixed paraffin embedded tissues. The results were correlated each other and with clinicopathologic parameters using Chi Square statistics.
Results: The rate of hypermethylation of GSTP1 was 28%. Of which, 8 cases showed double-band signal indicating either heterozygous tumors or contamination with normal tissues. Another case was found in a noninvasive carcinoma. Of the remaining 19 cases with hypermethylation status, 12.5% contradictorily revealed protein expressions. The correlation of the results from MSP and IHC methods yielding the Chi Square test, p-value= 0.04. The correlations with clinicopathological parameters showed that progesterone receptor correlated with MSP-methylation status (p-value= 0.05) and estrogen receptor correlated with GSPTP1 expression (p-value= 0.001). The other parameters - tumor size, tumor grade, lymph node status, HER2-IHC score, Ki67 index were not correlated.
Conclusion: The results between hypermethylation of GSTP1 by MSP and GSTP1 expression by IHC correlate, however, some unclear technical problems exist.
Keywords: Breast cancer, Glutathione S-transferase, promoter methylation, GSTP1
INTRODUCTION
Glutathione S-transferase class P1 (GSTP1) is one of the xenobiotic metabolizing genes. The encoded enzyme helps catalyzing the conjugation of glutathione to toxic compound that enhances excretion of the unwanted products [1]. This mechanism is important for host defense to carcinogenesis. Hy-permethylation state is linked to precursor of prostatic adenocarcinoma [2]. The reported rates of promoter methylation in breast cancer vary from 10% to 30% [3,4,5,6,7,8]. The event has claimed to be predictive of drug treatment [9]. With unexplained reason, there were 12% of hypermethylation cases that expressed GSTP1 protein [8]. This study is aimed to investigate whether the finding is natural variation or laboratory factors.
MATERIAL AND METHOD
Studied samples
The studied samples were frozen breast tumor tissues that had been collected in the tissue bank (-80°C) of Department of Pathology, Faculty of Medicine, Chulalongkorn University during 2006-2011. Gross malignant tissues were taken with a maximum dimension of 0.5 cm each case and kept in the refrigerator until experiment. The corresponding paraffin blocks of each tumor (case) were retrieved and one representative block was selected for immunohistochemistry study while the fresh frozen tissues were used for methylation specific PCR (MSP) assay. This study protocol has been approved by the faculty IRB.
MSP assay
Genomic DNA was extracted from the frozen tumor tissues using Qiaamp DNA FFPE tissue Kit (Qiagen, USA). When tissues thawed, they were chopped up and placed in a microcentrifuge tube that contained 180 pl of ALT buffer and 20 pl of proteinase K. The solutions were incubated at 56 °C overnight and at 90 °C for one hour. After that 200 pl ALT buffer and absolute alcohol were poured subsequently. The solutions were put into the QIAamp MinElute columns, centrifuged at 14,000 rpm for one minute, then the first rinse by 500 pl AW1 buffer in one minute centrifugation and the second rinse by 500 pl AW2 buffer in four-minute centrifugation. The QIAamp MinElute columns, placed in microcentrifuge tubes were filled with100 pl ATE, and incubated at room temperature for 10-20 minutes. After centrifuge at 14,000 rpm for one minute, the QIAamp MinElute columns were put away, leaving the DNA extracts solutions in the tubes that were kept at -20°C until use. The quantity of DNA was tested by Nanodrop Spectrophotometer (Thermo Scientific Corp.).
Sodium bisulfite treatment was done by using Bisulfite EZ DNA methylation gold kit (Zymo Research, Orenge, CA). Briefly CT conversion reagent was prepared by adding 900 pl distill water, 300 pl M-dilution buffer and 50 pl M-dis-solving buffer. One tube of CT conversion was used for 10 samples. Add 130 pl of the CT conversion reagent to 20 pl of DNA sample in a PCR tube. The tubes were placed in Thermal Cycler, incubated at 98°C for 10 minutes and 64°C for 2.5 hours. The working DNA extracts could be stored at 4°C up to 20 hours according to the manufacture. There were two pairs of primers. The primer sequences for the unmethyllated reaction were 5’-GAT GTT TGG GGT GTA GTG GTT GTT-3’ (forward primer) and 5-CCA CCC CAA TAC TAA ATC ACA ACA-3’ (reverse primer). The primer sequences for the methylated reaction were 5’-TTC GGG GTG TAG CGG TCG TC-3’ (forward primer) and 5’-GCC CCA ATA CTA AAT CAC GAC G-3’ (reverse primer) [8,9], PCR amplifications were done under the following conditions: 1 cycle at 95°C for 7 min, and 40 cycles at 95°C for 1 min, 61°C for 1 min and 72°C for 1 min, final extension at 72 oC for 7 min. PCR reactions were analyzed on 8% polyacrylamide gel electrophoresis, stained with Gelstar nucleic acid stain and visualized under UV illumination. The methyllation band was present at 91 base-pairs and unmethyllation band at 97 base pairs.
Immunohistochemical assay
Two-pm sections were cut and processed according to the standard immunohistochemistry method. The anti-GSTP1 antibody produced in rabbit (Sigma Corp, USA) was used. The optimal dilution had been calibrated at 1:800. All the im-munohistochemistry steps were done by Autoim-munostainer (Ventana, Roche).
GSPT1 expression was considered to be positive when >10% of tumor cells exhibited cytoplasmic or nuclear staining [10]. Two pathologists independently interpret the staining. If any cases were not in agreement, the slides would be looked again together to have a consensus result.
Clinicopathologic parameters
The following parameters were retrieved from the faculty data base - age, tumor size, tumor grade, immunohistochemistry results for ER, PR, HER2, p53 and Ki67, and lymph node status.
Statistics.
The relationship between GSTP1 methyla-tion status and immunohistochemical expression results was examined by Chi square. The test was also used for correlation assessments for both GSTP1 methylation status and protein expression status with clinicopathological parameters.
RESULTS
Of the 100 cases (samples), MSP assays show 72 unmethylation and 20 methylation cases. The determinate cases have single-band signals in either unmethylation or methylation (figure 1). There are 8 cases showing double-band signals, both unmethylation and methylation are called undetermined cases (figure 2).
The immunohistochemistry study reveals that 96 cases out of the 100 samples have sufficient invasive carcinoma. Of these, 52 cases are positive and 44 cases are negative according to the interpretation criteria above (figure 3). It is remarkable that the pre-existing normal ductal cells are strongly positive and can be used as internal control (figure 4). On the other hand, tumor cells in some cases display composite areas of positive and negative staining (figure 5).
Upon investigation of the relationship of results between MSP and IHC methods, twelve cases are excluded. Four cases are deleted because the tissue sections consist mostly or exclusively of non-invasive carcinoma components (MSP results show one methylation and three unmethylation cases). The other eight discarded cases are due to two-band signals by MSP (four cases show positive and four negative IHC results). The analysis of the 88 cases reveals a statistically negative relationship. (Table 1)
The clinicopathological parameters have been correlated with both MSP status and IHC status separately. There are no significant relationships except for ER and PgR. (Tables 2, 3)
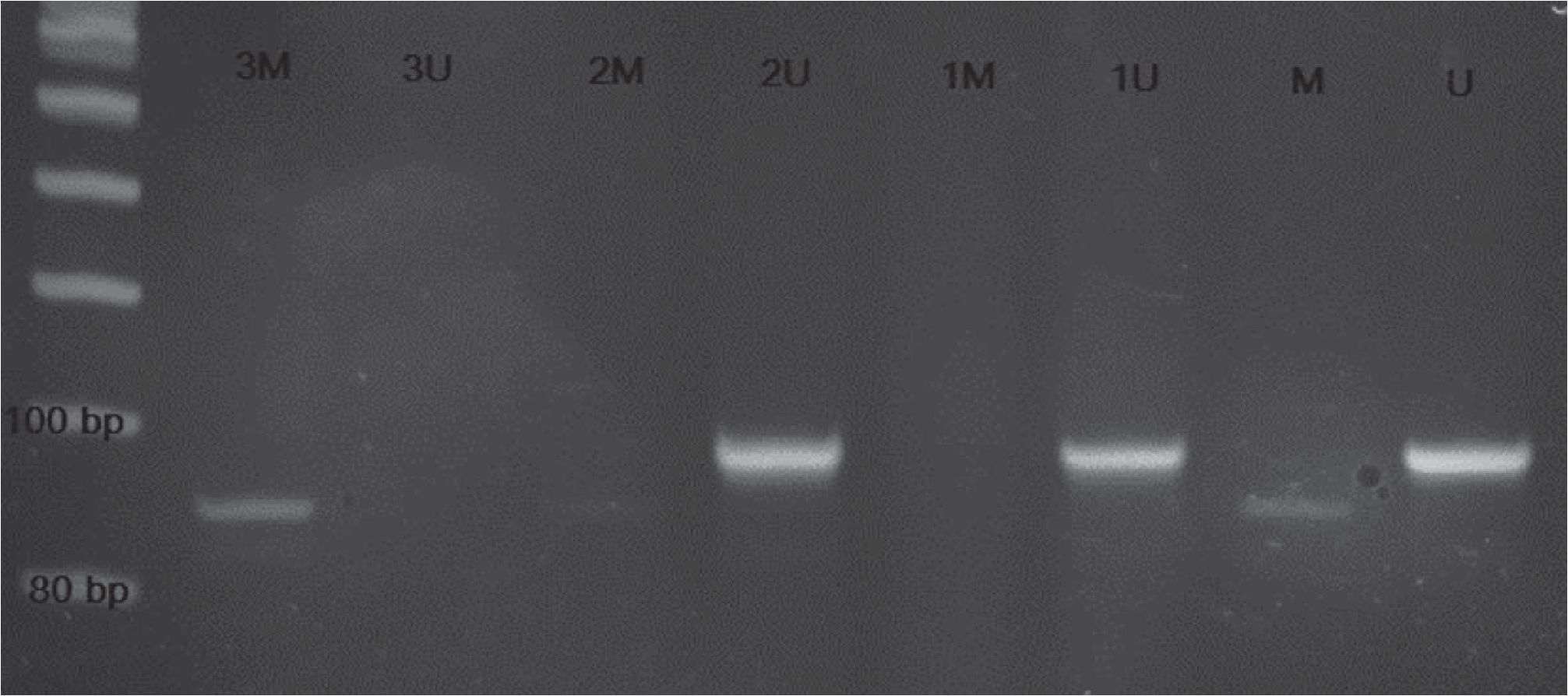
Figure 1. DNA bands of representative cases showing unmethylation and methylation. Cases 1 and 2 comprise single band signal of unmethylation (lanes labeled as 1U and 2U respectively) whereas case 3 reveals single band signal of methylation (lane labeled as 3M).

Figure 2. DNA bands of representative cases showing undetermined and unmethylation. Cases 4 and 5 comprise double band signal of methylation and unmethylation (lanes labeled as 4M/ 4U and 5M/5U respectively) whereas case 6 reveals single band signal of unmethylation (lane labeled as 6U).
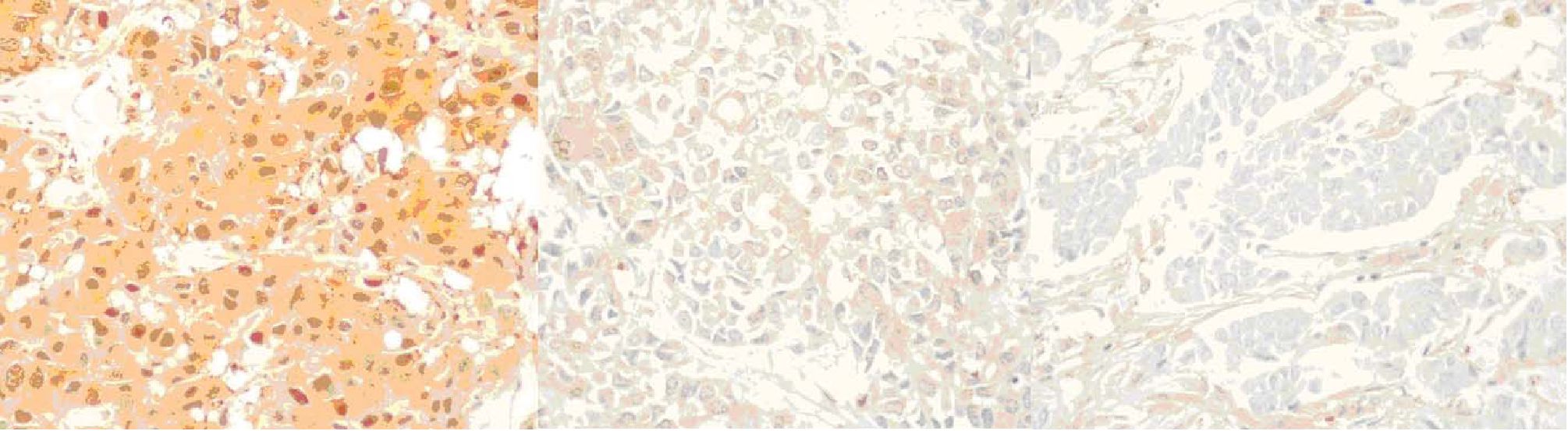
Figure 3. GSTP1 expressions in representative cases showing strongly positive, weakly positive and negative immunohistochemistry staining. (x 400 original magnification)
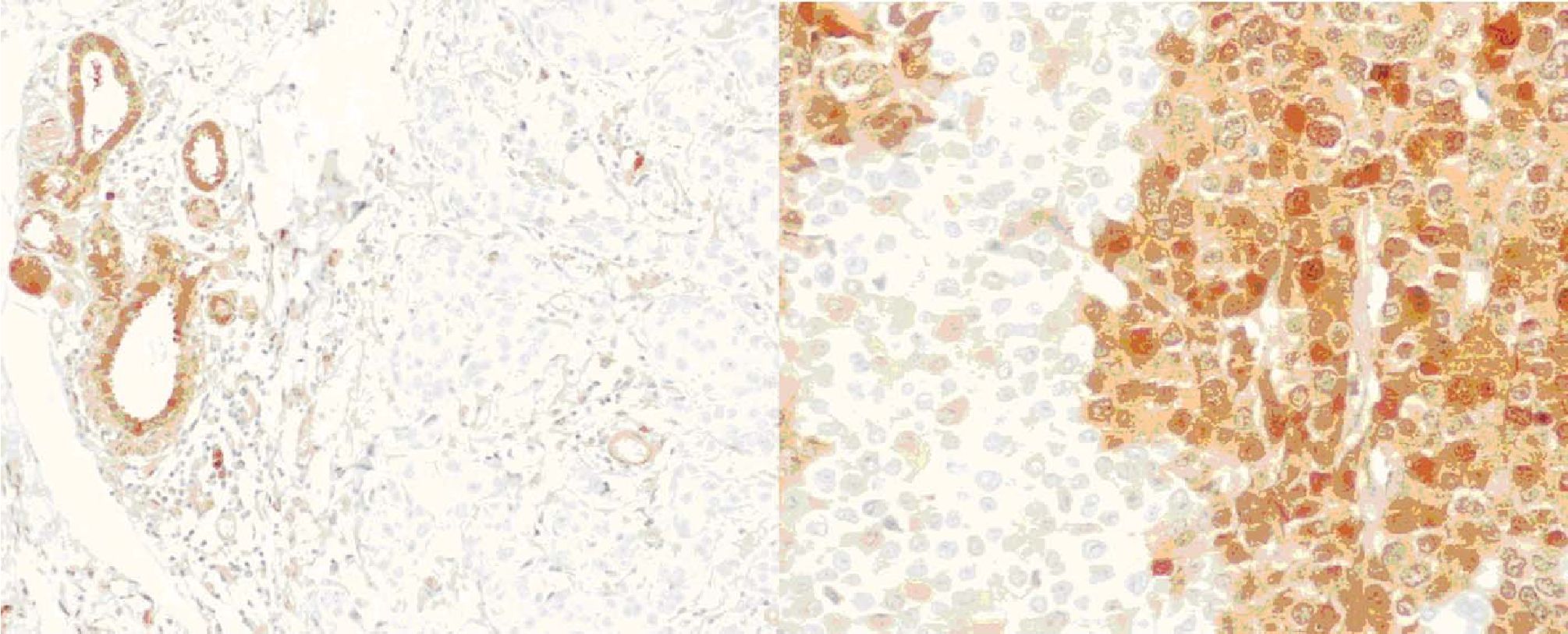
Figure 4. GSTP1 expressions showing one case with negative staining of tumor cells and positive staining of normal ductal cells and the other case with composite areas of weak and strong staining of tumor cells. (x 200, x 400 respectively by original magnification)
Table 1. Relationship between MSP assays and IHC expressions
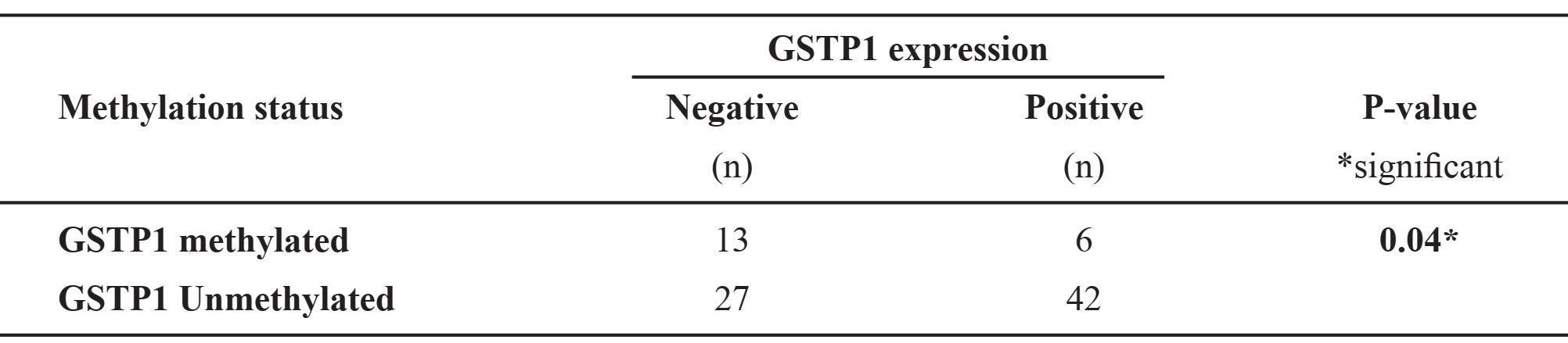

Table 2. Correlation of Methylation status and clinicopathologic parameters
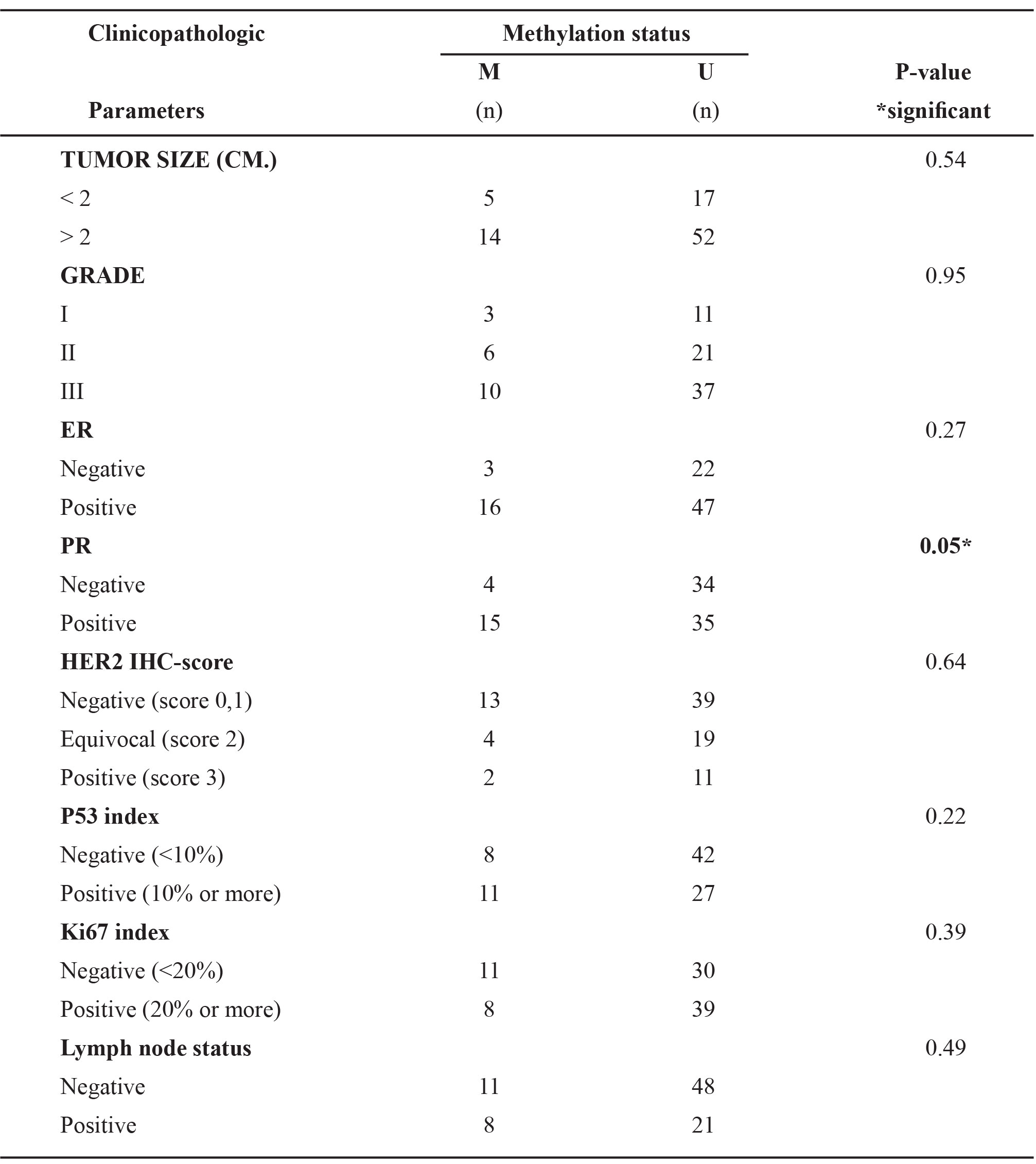
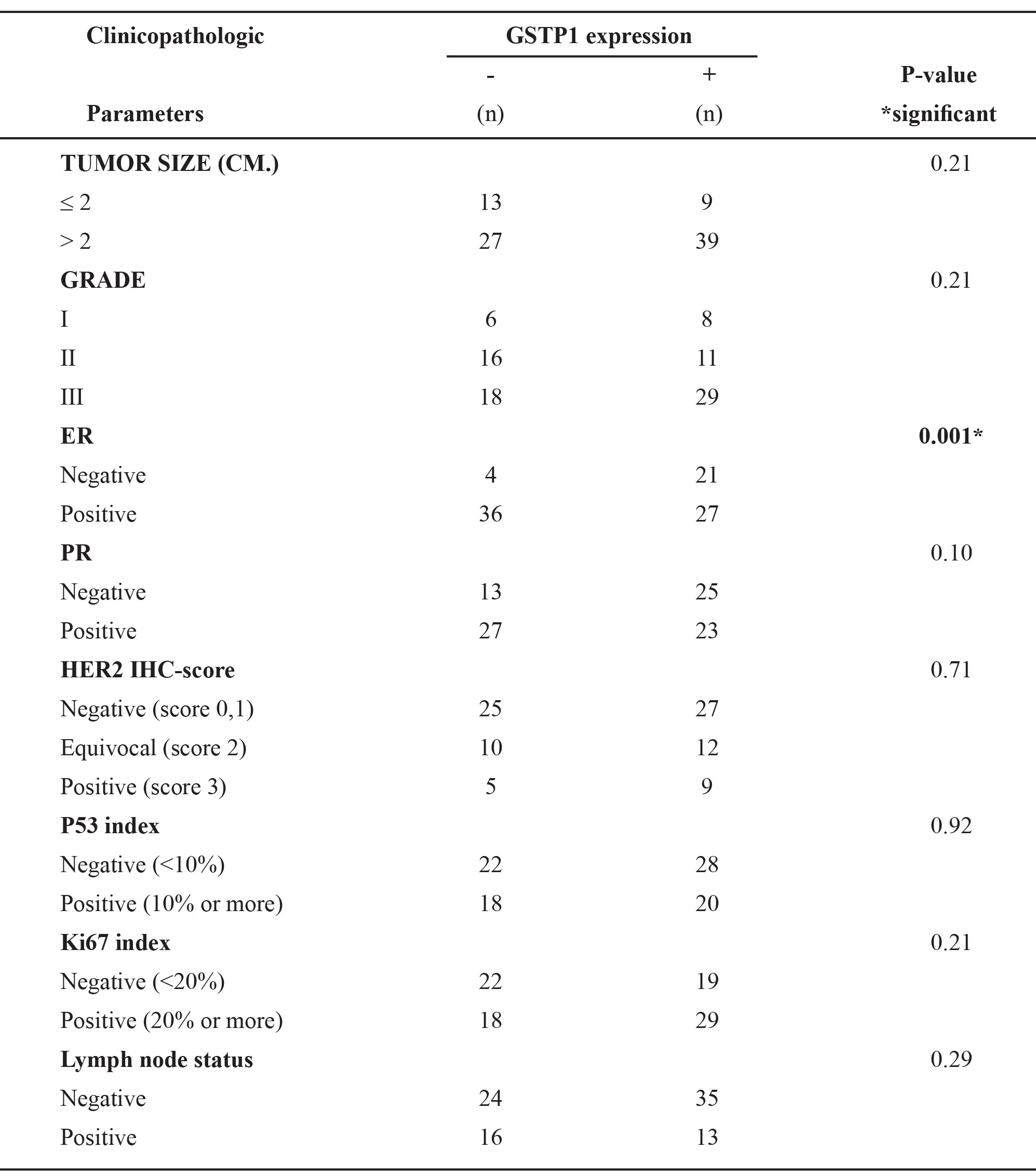
Table 3. Correlation between immunohistochemistry status of GSTP1 and clinicopathologic parameters
DISCUSSION
There are many methods of studying meth-ylation events. Methylation specific PCR (MSP) is the most widely used technique owing to high sensitivity and specificity. The method does not require large DNA amount [11]. Nevertheless, MSP assay gives less resolution compared to Bisulfite Sequencing technique. The latter discloses base-by-base. Immunohistochemistry staining (IHC) is practical and a familiar technique for pathologists. It allows histomorphologic correlation. Theoretically, the results from these two methods should correlate. The hypermethylation cases should not express the protein. The finding of 12% of hypermethylation cases expressing GSTP1 in the previous study interested the authors whether this is an unclear natural variation or laboratory problem. We speculated that in the natural variation, the similar finding would be found. In addition, this repeated experiment may indicate some laboratory problems.
Our study reveals the prevalence of hypermethylation of GSTP1 is 28%. If 9 cases were excluded, the 22% prevalence is still in the previous reporting range [3,4,5,6,7,8]. Our result is also in agreement with the previous study that about 12.5% of the hypermethylation cases show positive im-munostaining for GSTP1. The prior authors did not comment on this occurrence; we speculate that the primer probes of methylation and unmethylation may not cover the whole active transcription bases. Other methods are needed to verify this technical problem.
Regarding DNA extraction, fresh tissue samples are tremendous better than formalin fixed paraffin embedded tissues [7]. In this study, the quantity of DNA extracts was fruitful in all cases. The limitations were that the collected fresh frozen tissue samples were small, not enough for histo-morphological confirmation and not available for the immunohistochemistry study for protein expression.
Bisulfite treatment seems to be one of the critical steps. It will convert unmethylated cytosine into uracil. The duration and volume of substrates are essential for complete reactions. In the primer design, the MSP used two separate sets of methylation probe and unmethylation probe. In cases of hy-permethylation or unmethylation, single band signal on the hypermethylation and unmethylation are illuminated respectively. The presence of doubleband signal is problematic. There are eight cases in this study but in other previous publications this finding was vague. We suggest two possibilities. It might be caused by heterozygous tumor or contamination of the normal breast tissue. The small chance of incomplete bisulfite-reaction is arguable [12]. In these eight cases, four cases are protein positive and four others are negative. The cause was not simply sorted. We therefore treated the eight cases as problematic cases and data were excluded on the analysis of the relationships with immunohistochemistry as well as clinicopathological parameters.
IHC results were clear. Inter-observer correlation was obtained in all cases. Because normal duct cells are obviously positive and no information about non-invasive carcinoma should be assessed or not, the authors excluded four cases that contained mainly non-invasive carcinoma component from the relationship analysis.
On the correlational study, the results between MSP and IHC were statistically significant, however, the eight cases of double-band signal that were excluded are cautious. Further study to verification the properness of primers is needed. In addition, why some cases (12.5%) of hypermethyla-tion status expressed protein also requires further explanatory experiment.
The correlations of the MSP and IHC with clinicopathological parameters are not well-established. Like in the other papers that correlations were diverse. It seems that some laboratory factors have to fix before clinical research based on MSP study can go further.
In conclusion, the results between hyper-methylation of GSTP1 by MSP and GSTP1 expression by IHC correlate in some extent, however, there might be unclear technical problems regarding MSP that need further verification.
REFERENCES
1 Coles BF, Kadlubar FF. Detoxification of electrophilic compounds by glutathione S-transferase catalysis: determinant of individual response to chemical carcinogen and chemotherapeutic drugs. Biofactors 2003; 17: 115-130.
2 Yennasubramanian S, Kowalski J, Gonzalgo ML, et al. Hypermethylation of CpG islands in primary and metastatic human prostate cancer. Cancer Res 2004; 64: 1975-1986.
3 Parrella P, Poeta ML, Gallo AP, Prencipe M, Scintu M, Apicella A, Rossiello R, Liguoro G, Seripa D, Gravina C, Rabiti C, Rinaldi M, Nicol T, Tommasi S, Paradiso A, Schittulli F, Altomare V, Fazio VM. Nonrandom distribution of aberrant promoter methylation of cancer-related genes in sporadic breast tumors. Clin Cancer Res 2004; 10: 5349-5354.
4 Shinozaki M, Hoon DS, Giulia AE, Hansen NM, Wang HJ, Turner R, Taback B. Distinct hypermethylation profile of primary breast cancer is associated with sentinel lymph node metastasis. Clin Cancer Res 2005; 11: 21562162.
5 Esteller M, Corn PG, Urena JM, Gabrielson E, Baylin SB, Herman JG. Inactivation of glutathione S-transferase P1 gene by promoter hypermethylation in human neoplasia. Cancer Res 1998; 58: 4515-4518.
6 Lee JS. GSTP1 promoter hypermethylation is an early event in breast carcinogenesis. Virchows Arch 2007; 450: 637-642.
7 Pongtheerat T, Pakdeethai S, Purisa W, Chari-yalertsak S, Petmitr S. Promoter Methylation and Genetic Polymorphism of Glutathione S-Transferase P1 Gene (GSTP1) in Thai Breast cancer Patients. Asian Pacific J Cancer Prev 2011; 12: 2731-2734.
8 Arai T, Miyoshi Y, Kim SJ, Taguchi T, Tamaki Y, Noguchi S. Association of GSTP1 CpG islands hypermethylation with poor prognosis in human breast cercers. Breast Cancer Res Treat 2006; 100: 169-176.
9 Arai T, Miyoshi Y, Kim SJ, Akazawa K, Maruyama N, Taguchi T, Tamaki Y, Noguchi S. Association of GSTP1 expression with resistance to docetaxel and paclitaxel in human breast cancers. Eur J Surg Oncol 2008; 34: 734-738.
10 Huang J, Tan PH, Thiyagarajan J, Bay BH. Prognostic significance of glutathione S-transferase-pi in invasive breast cancer. Mod Pathol 2003; 16: 558-565.
11 Fraga MF, Esteller M. DNA Methylation: A profile of Methods and Applications. BioTechniques 2002; 33: 632-649.
12 Thomassin H, Oakeley EJ, Grange T. Identification of 5’-methylcytosine in complex genomes. Methods 1999; 19: 465-475.


