Angioimmunoblastic T-Cell Lymphoma: an Epstein-Barr Virus-Associated, Bilineage Lymphoid Neoplasm
Surapong Sorcharoen, MD, Supaporn Suwiwat, PhD.* Winyou Mitarnun, MD.
Department of Pathology, Faculty of Medicine Prince of Songkla University, Hat-Yai, THAILAND 90110
Correspondence Winyou Mitarnun, MD
E-mail: miwinyou@medicine.psu.ac.th Tel: +66 074 45 1551 Fax: +66 074 212908
Received 13 Nevember 2013; Accepted 2 February 2014
Abstract
In this study, 50 cases of angioimmunoblastic T-cell lymphoma (AITL) were investigated for histological patterns, their association with Epstein-Barr virus (EBV) infection, B cell and T cell clonalities, and the clinical outcomes. Three histological patterns were identified: the classical pattern (34 cases), large-cell rich pattern (14 cases), and interfollicular pattern (2 cases). The median survival time of patients from this group was 36 months. There was no significant difference in survival between patients from the classical pattern and the large-cell rich pattern. EBV-encoded early RNAs (EBERs) in tumor cells were identified in 46% of the cases. The expression of EBERs had no correlation with the histological patterns. We detected T-cell receptor-gamma chain (TCR-γ) T-cell clones in 70% of the cases, and immunoglobulin heavy chain (IgH) B-cell clones in 32% of the cases. Bilineage monoclonal proliferation of both T and B cells were detected in 28% of the cases. Clonal B cell expansion had no correlation with number of B cells in the lymph node and the status of EBERs in the tumor cells. Based on the findings from molecular study, we believe that AITL is a bilineage lymphoid neoplasm and may contribute to the pathogenesis of composite lymphomas.
Keywords: Angioimmunoblastic T-cell lymphoma, Epstein-Barr virus, EBV-encoded early RNA, Clonality, T-cell receptor (TCR), immunoglobulin heavy chain (IgH), Composite lymphoma
Running Title: Angioimmunoblastic T-cell lymphoma
INTRODUCTION
Angioimmunoblastic T-cell lymphoma (AITL) is a systemic disease of peripheral (mature) T cell neoplasm found mainly in the lymph nodes but may also occur in the spleen, bone marrow, liver, and skin1-3. Histologically, the lymph nodes show partial or totally obliteration of the normal architecture by a polymorphic infiltrate lymphocytes, plasma cells, eosinophils, histiocytes and a proliferation of venules and follicular dendritic cells (FDCs)1,4. In many instances the histopathological features of malignancy are not readily identifiable. This disease was first described as an atypical reactive process, however, molecular studies have consistently shown clonal abnormalities, with monoclonal T cell proliferation in most cases and monoclonal B cell proliferation in some cases[3]. In Western countries, AITL is one of the more common specific subtypes of the peripheral T-cell lymphoma, accounting for approximately 15-20% of peripheral T-cell lymphoma cases. The primary site of this disease is the lymph nodes and it occurs mainly in the middle-aged and elderly (median age = 64 years) with an equal incidence in both sexes. In Thailand, the incidence of AITL is about 67% of nodal peripheral T-cell lymphoma cases, and occurs in every age group (median age = 51 years) with a male to female ratio of about 2:15.
AITL is highly associated with latent Epstein-Barr virus (EBV) infection. The presence of EBV genomes, EBV-encoded small nuclear RNAs (EBERs) in tumor cells is detected in most of AITL cases5-7. In one study, in about 26% of cases, EBERs were detected in both B and T cells, the majority of which were B cells of immunoblastic morphology7. In a study from Thailand5, EBERs-positive tumor cells were detected in 50% of AITL cases, and in most cases were detected in both background lymphocytes (T and B cells) and im-munoblasts. The presence of EBV genomes in tumor cells had no correlation with the clinical outcomes6.
Studies have found that T cell receptor (TCR) genes showed clonal rearrangement in 60% to 100% of cases8,9, and clonal immunoglobulin heavy chain (IgH) gene rearrangements were found in 8% to 40% of cases10,11. Composite lymphomas which composed of EBV-associated AITL and EBV-associated diffuse large B-cell lymphoma and EBV-associated diffuse large B-cell lymphoma arising in the preexisting EBV-associated AITL have also been reported12,13.
The aim of this study was to investigate the clonality of the T and B cells in EBV-positive and EBV-negative AITL cases in Thailand and to correlate these findings with histological patterns and clinical outcomes.
MATERIALS AND METHODS
Patient Samples
All ofthe cases were from the Department of Pathology, Faculty of Medicine, Prince of Songkla University, Songkhla, Thailand. Consecutive cases from 1990 through 2007 were examined. There were 50 cases of AITL in the lymph nodes. The diagnosis of AITL was made according to the WHO classification!. Three patterns of AITL were recognized: classical, large-cell rich and interfollicular14-16.
Morphological criteria of AITL subtypes
Classical pattern (Fig.1): This pattern showed partial or diffuse effacement of the nodal architecture, vascular proliferation with prominent abori-zation of high endothelial venules, extrafollicular meshwork of FDCs, an atypical population of CD3+ T cells, and large CD20+ B cells. The percentage of large cell was less than 10% of the total cell population. The cellular infiltration was admixed with variable numbers of eosinophils, plasma cells, histiocytes and occasionally Reed-Sternberg-like B cells and granulomatous formations were seen.
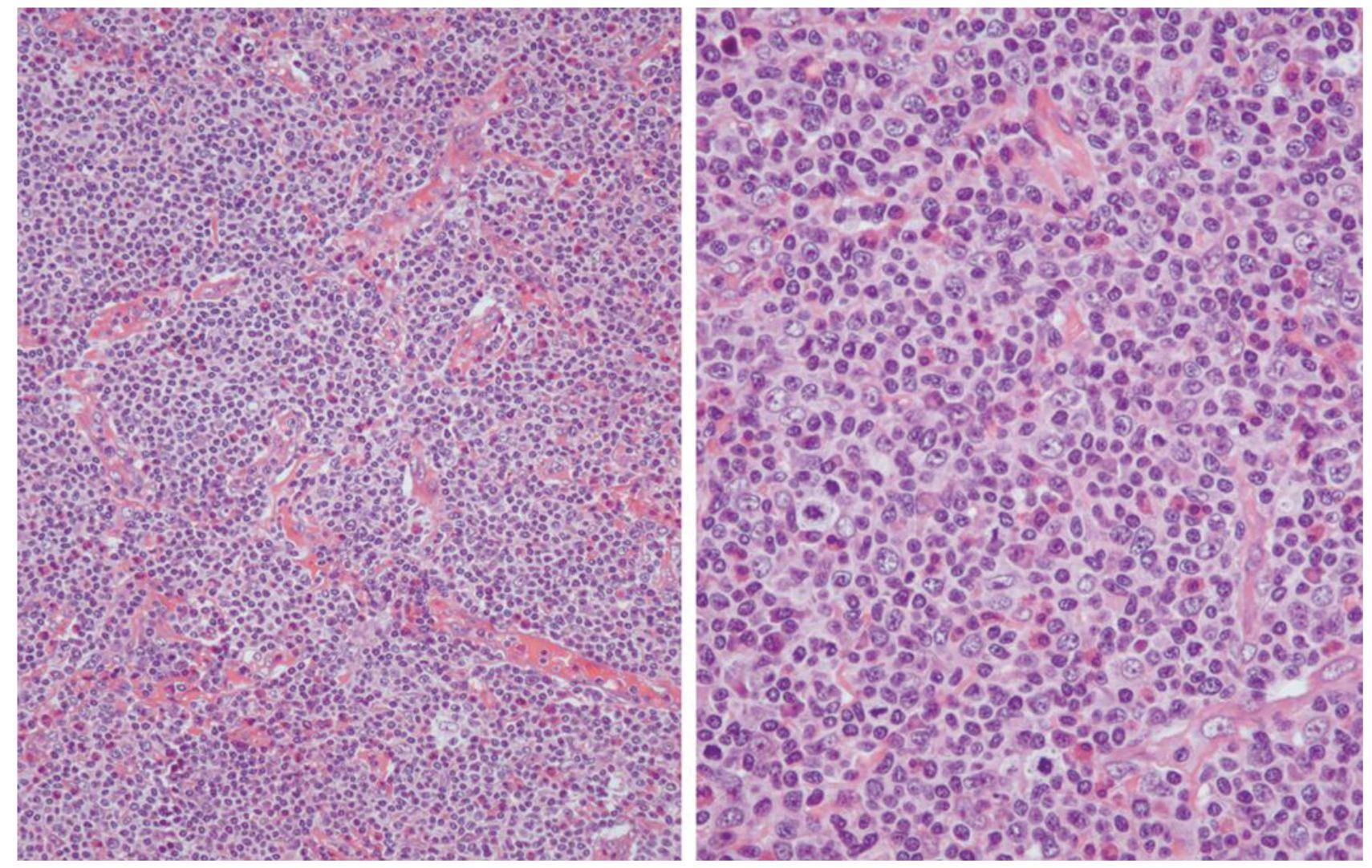
Figure 1 Angioimmunoblastic T-cell lymphoma, classical pattern.
Large-cell rich pattern (Fig.2): This subtype of AITL had histopathological changes similar to the classical pattern except for an increase in number of large cells. The large cell population (B and/or T cells) was more than 10% of the total cell population.
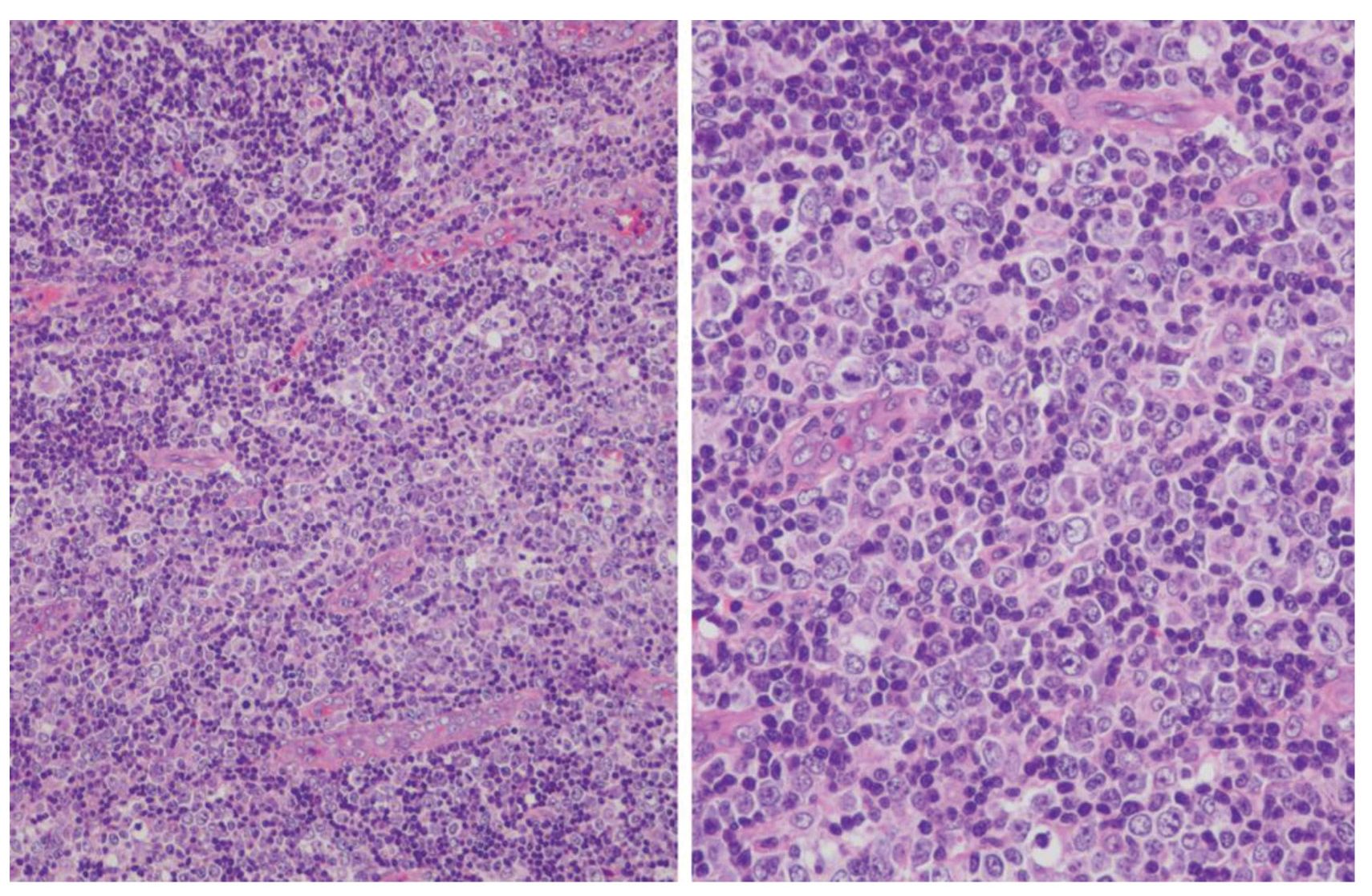
Figure 2 Angioimmunoblastic T-cell lymphoma, large-cell rich pattern.
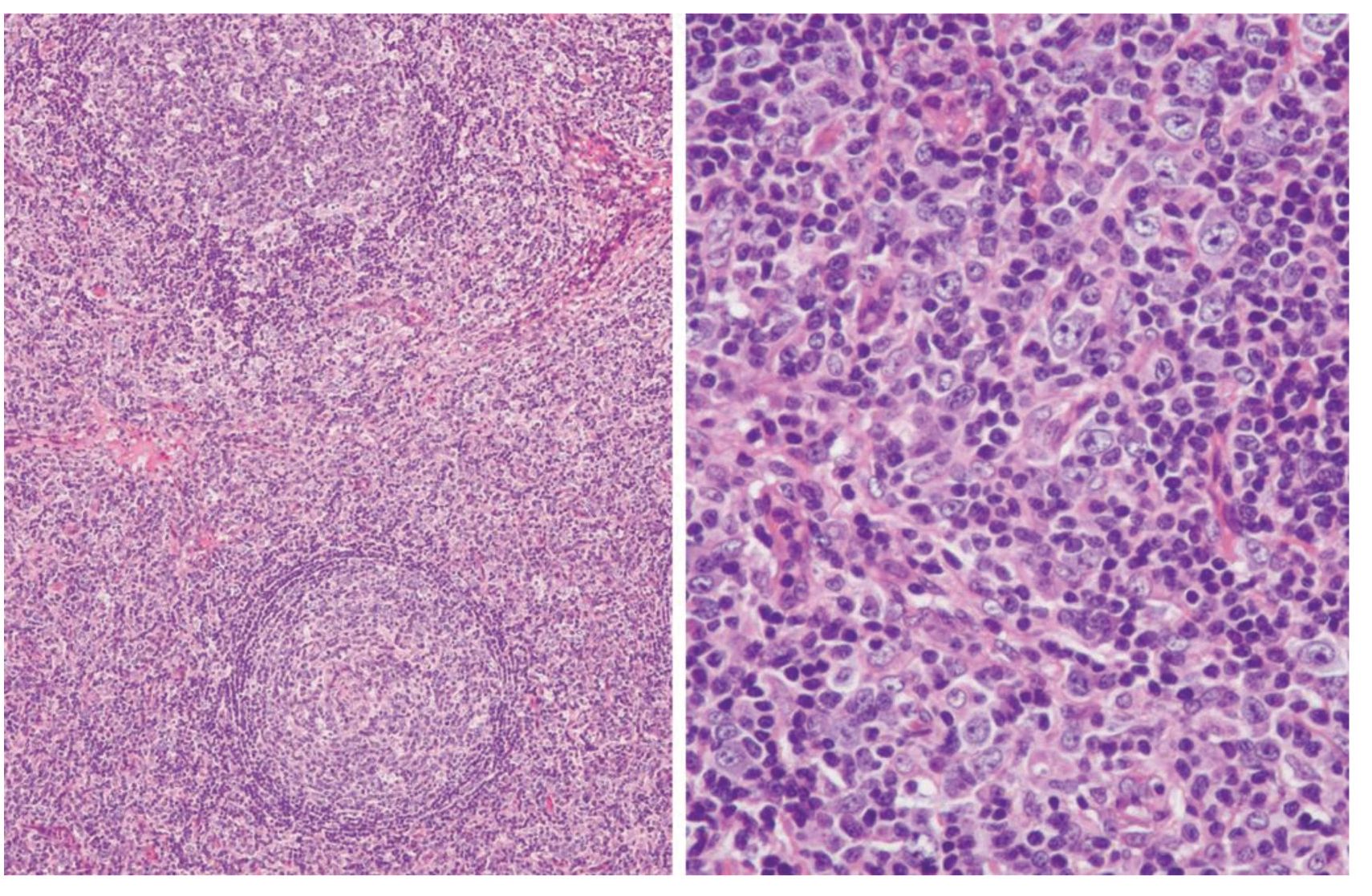
Figure 3 Angioimmunoblastic T-cell lymphoma, mterfollicular pattern.
Interfollicular pattern (Fig.3): This pattern had paracortical expansion with hyperplastic germinal centers. The histopathology in this subtype consisted of the combination of hyperplastic germinal centers along with angioimmunoblastic reaction.
Immunohistochemistry
Immunohistochemical staining for the leukocyte common antigen (LCA), CD3, CD5, CD10, CD20, CD23 was performed on formalin-fixed, paraffin-embedded tissues using the following antibodies: monoclonal mouse antihuman LCA (CD45) (Dako Cytomation, Demmark), monoclonal mouse antihuman CD3 (Novocastra, UK), monoclonal mouse antihuman CD5 (Novocastra, UK), monoclonal mouse antihuman CD10 (Novocastra, UK), monoclonal mouse antihuman CD20 (Zymed Labaratories, USA) and monoclonal mouse antihuman CD23 (Zymed Laboratories, USA).
In situ hybridization study for EBV
Detection of EBV RNA expression was carried out using fluorescein-conjugated EBV oligonucleotides complementary to portions of the EBV-encoded early RNA transcripts (EBERs) (EBER1/2, Novocastra, UK) that were actively transcribed in latently infected cells, as previously described17. Appropriate positive and negative controls were run in every batch tested. Few positive cells or positive less than 5% of the total tumor cell population were considered to be negative5.
DNA extraction from the lymph nodes for PCR analysis
DNA extraction was performed using a QIAamp DNA Mini Kit (QIAGEN, Germany). Three to five pieces of tissue section (3-5 pm) were taken from paraffin blocks, deparaffinized, resuspended in 180 pL of buffer ATL, digested with 50 pL of proteinase K, mixed by vortexing and incubated at 55°C until the tissue was completely lysed. Additional steps followed the tissue protocol outlined in the manufacturer’s manual. The presence of DNA was confirmed by agarose-gel electrophoresis.
Clonality studies of the tumor cells
The TCR-γ chain gene was amplified using the method as previously described[18]. We used five separate reactions with the following oligonucleotide primers combination: VyI+VynI/IV+Jy1/2 (product sizes 70-95 base pairs), VyI+VyIII/IV+JPy1/2 (product size 80-100 base pairs), VyII+Jy1/2 (product sizes 150-180 base pairs) VyII+JPy1/2 (product sizes 160-190 base pairs), and VyI+JPy (product sizes 70-95 base pairs). The sequences of oligonucleotide primers were listed in Table 1. For the monoclonal rearrangement, one or two clearly visible band/s were identified.
Determination of IgH gene rearrangement was performed by seminested PCR using FR1c-JH,
FR2a-JH and FR3a-JR primer amplification (Table1) [19]. Briefly, each 25 pl reactions mixture contained 5x PCR buffer, 2 mM MgCl2, 200 pM of each dNTP, 10 pmol of each primer, 1.25 U of Fast Start Taq DNA polymerase (Roche Diagnostics GmbH, Mannheim, Germany) and about 200 ng of DNA. Each PCR cycle consisted of 1 min at 93°C, 1 min at 54°C for FR1c and FR3a, and 50°C for FR2a, and 1 min at 72°C. For the first round of PCR amplification, the primers FR1c, FR2a, FR3a and LJH were used. For the second round of PCR, the primers FR2a and FR3a were used with inner primer VLJH. The first round of PCR consisted of 30 cycles and 1% of the PCR amplification product was used as a template for the second round of PCR consisting of 20 cycles. The PCR products were analyzed using 6% and 10% nondenaturing polyacrylamide gel electrophoresis, stained with ethidium bromide.
Table 1 Sequence of primers Primer

RESULTS
Clinical data
The patients consisted of 35 males and 15 females, (male: female=2.3:1), ranging in age from 17 years to 84 years; 5 patients (10%) were under 20 years of age. The median age was 56 years, and the mean (+SD) age was 50.7+19.3 years. Using the morphological criteria of the AITL subtypes, 34 patients were classified into the classical pattern (27 males, 7 females), 14 into the large-cell rich pattern (8 males, 6 females), and two female patients showed the interfollicular pattern. Kaplan-Meier survival estimates for patients from the classical pattern, large-cell rich pattern and all patterns combined are shown in Fig.4.
The median survival time of all patterns was 36 months. The median survival times of patients from the classical pattern and large-cell rich pattern were 24 months and 40 months, respectively. There was no significant difference in the survival profile between patients with the classical pattern and large-cell rich pattern (P=0.5464). The expression of EBERs in the tumor cells had no association with the disease outcome (P=0.5987).
Immunohistochemistry
Results of the immunohistochemistry are summarized in Table 2. The majority of lymphoid infiltrates were CD3+ cells in 72% of cases, and CD20+ cells in 14% of cases. Fourteen percent of the cases had an equal amount of CD3+ and CD20+ cellular infiltration. One patient showed dual staining of both CD3 and CD20 in nearly all tumor cells. Loss of CD5 expression on the surface of the T lymphocytes was detected in 6 of 48 cases (12.5%). CD10 expression was seen in 8 of 34 cases (23.5%) of the classical pattern and was not identified in the other two patterns. The fraction of CD10+ cells varied from around 5% to a maximum of 40% of the total lymphoid population. FDCs (CD23+ cells) were identified in 22 of the 47 cases (46.8%): 11 of the 31 cases in the classical pattern, 9 of the 14 cases in large-cell rich pattern and the 2 cases of the inter-follicular pattern. FDCs proliferation varied from minimal and limited to a few follicles, extended beyond the follicles into the paracortex, to the expansion into the interfollicular area and some FDCs were seen surrounding proliferating vessels. For the interfollicular pattern, the FDCs proliferation was limited to inside the hyperplastic follicles. In the large-cell rich pattern, the large cell populations were identified as CD3+ cells in 4 cases and CD20+ cells in 10 cases.
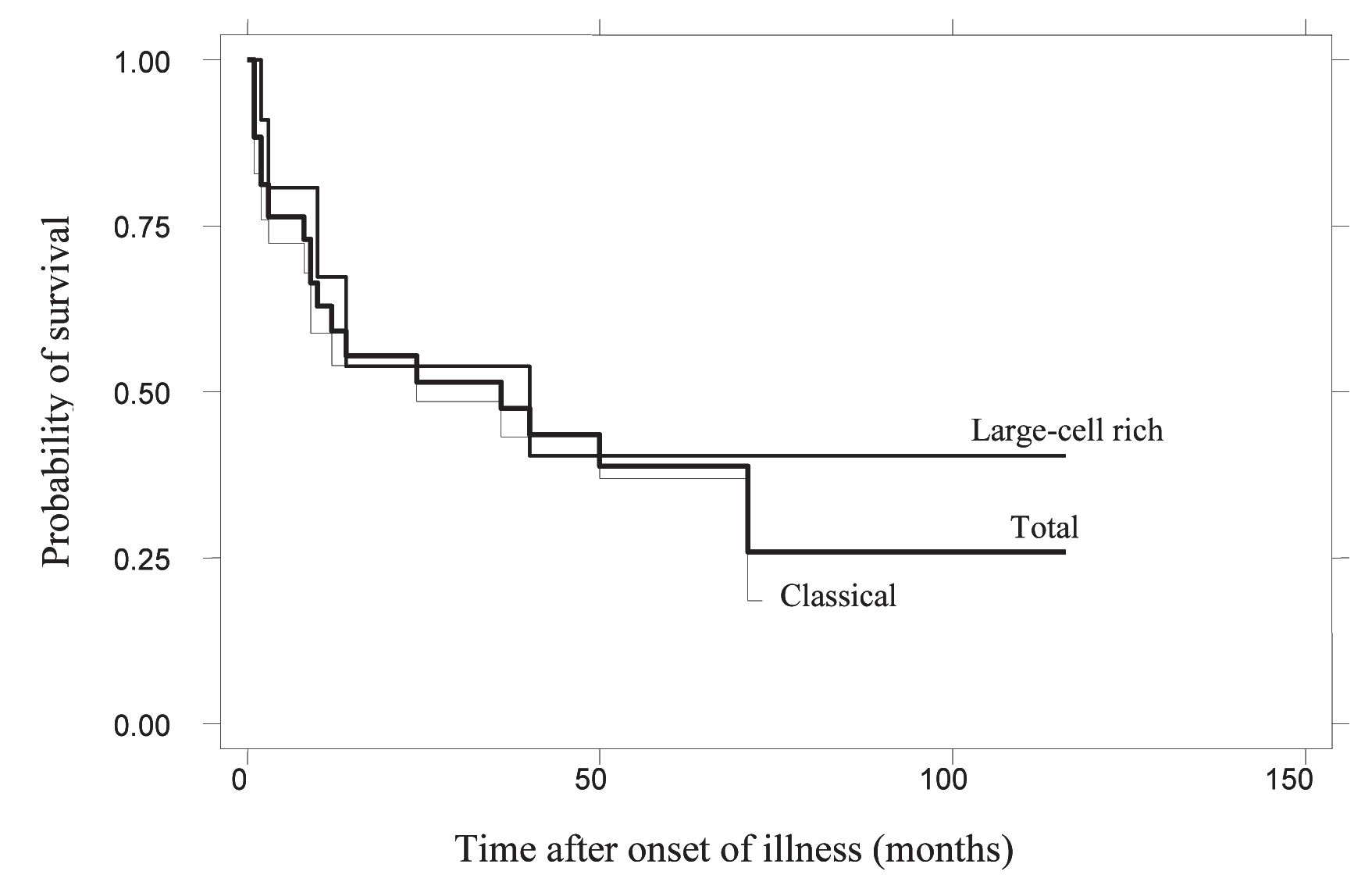
Figure 4 Probability of patient survival according to the patterns
Table 2 Immunohistochemistry of the AITL
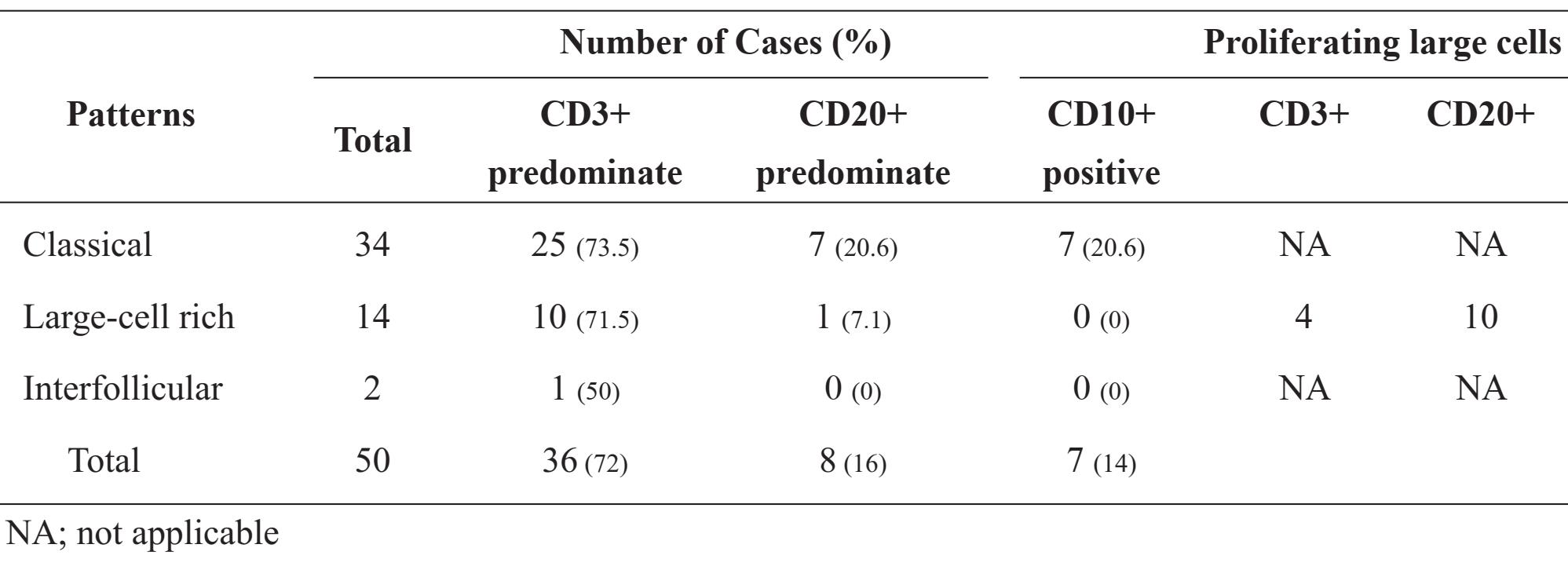
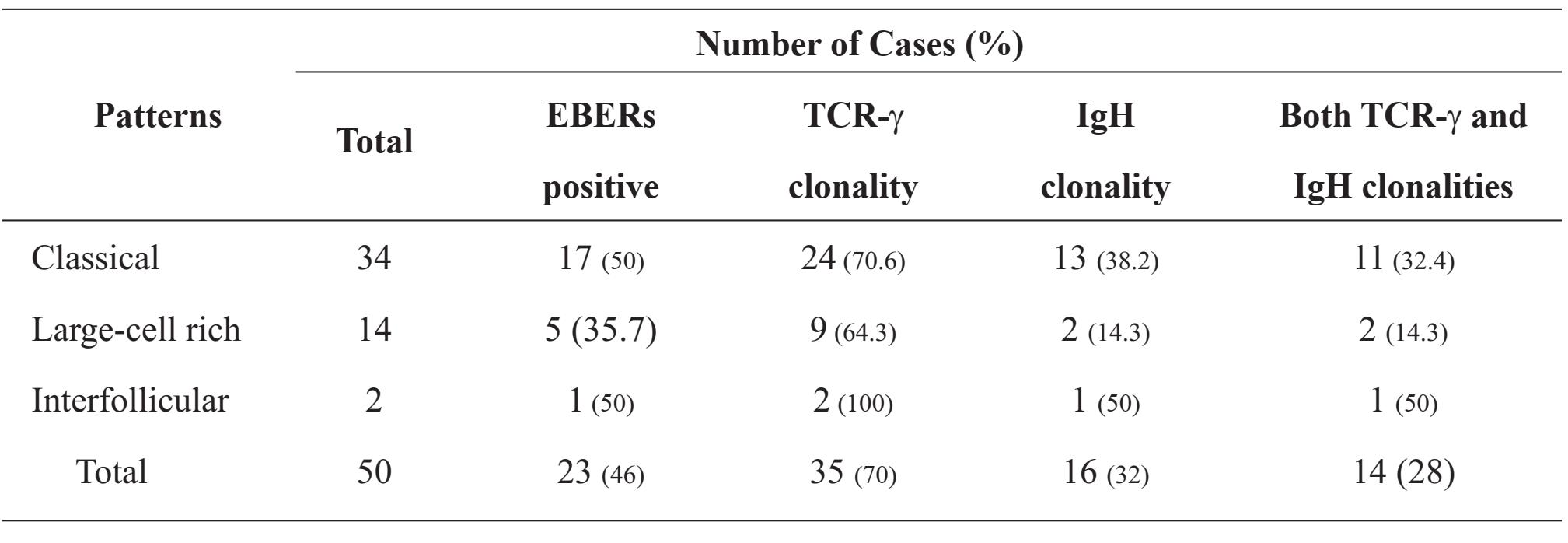
Table 3 EBV genomes and the clonality studies of AITL
EBV genomes and the clonality studies
The results of the EBV genomes and the clonality studies are summarized in Table 3. EBERs were identified by ISH in 23 of the 50 cases (46%). The frequencies of EBERs-positive in each of the histological patterns were as follows: classical pattern = 50%, large-cell rich pattern = 35.7%, and interfollicular pattern = 50%.
Using five sets of oligonucleotide primers for the TCR-γ gene, monoclonal band/s were detected in 35 of the 50 cases (70%): 70.6% in the classical pattern, 64.3% in the large-cell rich pattern, and two of the two cases (100%) in the interfollicular pattern. For the IgH gene, monoclonal band/s were detected in 16 of the 50 cases (32%): 38.2% in the classical pattern, 14.3% in the large-cell rich pattern, and one of the two cases (50%) in the interfollicular pattern.
Monoclonalities of both the TCR-γ gene and the IgH gene were identified in 14 of the 50 cases (28%), indicating bileneages monoclonal proliferations of T and B lymphocytes in a single lymph node. Of sixteen patients who showed monoclonality of the B lymphocytes, 14 also showed monoclonality of the T lymphocytes. Of seven patients with B cells predominate (more than 50% of the lymphoid infiltrates), four showed IgH clonal products. There was no correlation between B cell clonality and the presence of B cell proliferation (Pearson chi-squared, P=0.30). The presence of EBERs in tumor cells also had no correlation with the clonalities of either T or B cells (Pearson chi-squared, P=0.58 and 0.70, respectively).
Composite lymphomas
Three additional patients with nodal composite lymphomas of synchronous occurrence of AITL combined with peripheral T-cell lymphoma, unspecified (PTCL-NOS), anaplastic large-cell lymphoma, ALK-positive (ALCL-ALK+), and non-Hodgkin lymphoma, diffuse large B-cell (DLBCL) are shown in Fig.5-Fig.7, respectively. Figure 5 shows a lymph node from an 81-year-old female patient. Some portions of the lymph node were replaced by AITL and another portions were replaced by PTCL-NOS. EBERs were detected in both AITL and PTCL-NOS. Figure 6 shows a lymph node from a 23-year-old male patient. The lymph node showed simultaneous occurrence of admixed AITL and ALCL-ALK+, and the EBERs were negative in the entire lymph node. Figure 7 shows a lymph node from a 36-year-old male patient. Half of the lymph node showed AITL features and the other half was DLBCL. EBERs were detected only in the AITL.
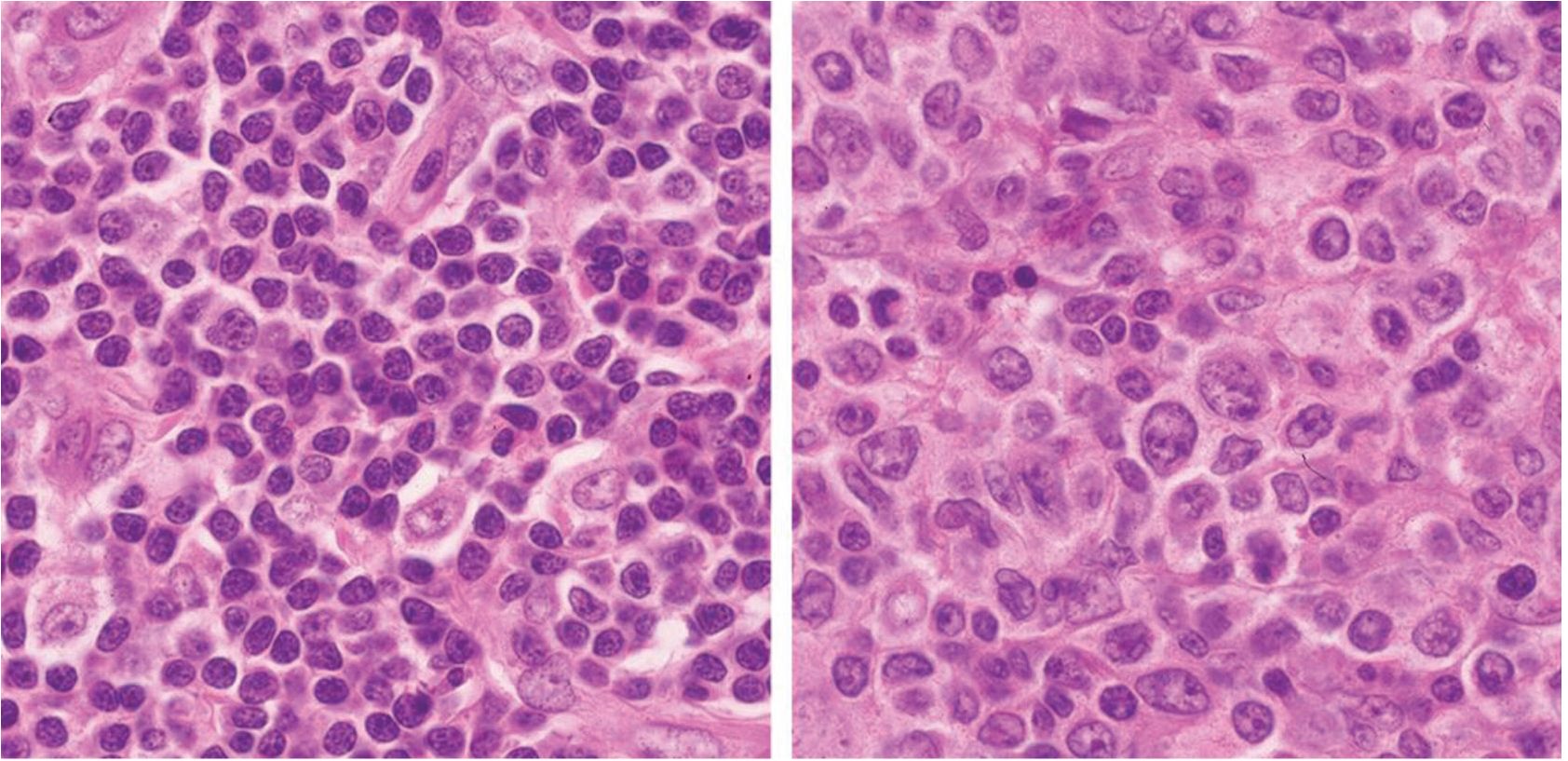
Figure 5 Composite lymphomas: angioimmunoblastic T-cell lymphoma (left) and peripheral T-cell lymphoma, NOS (right).
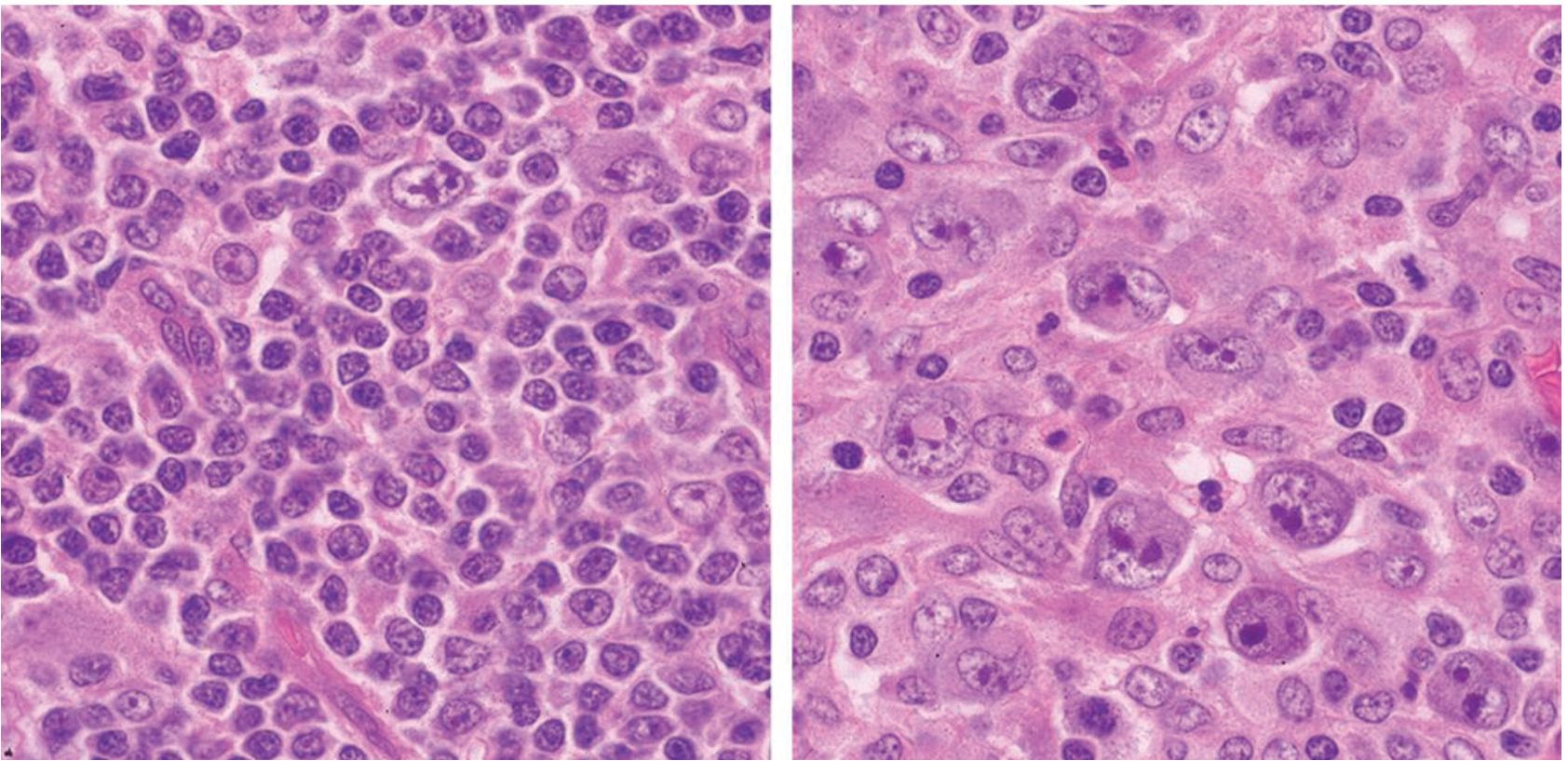
Figure 6 Composite lymphomas: angioimmunoblastic T-cell lymphoma (left) and anaplastic large cell lymphoma, ALK-positive (right).
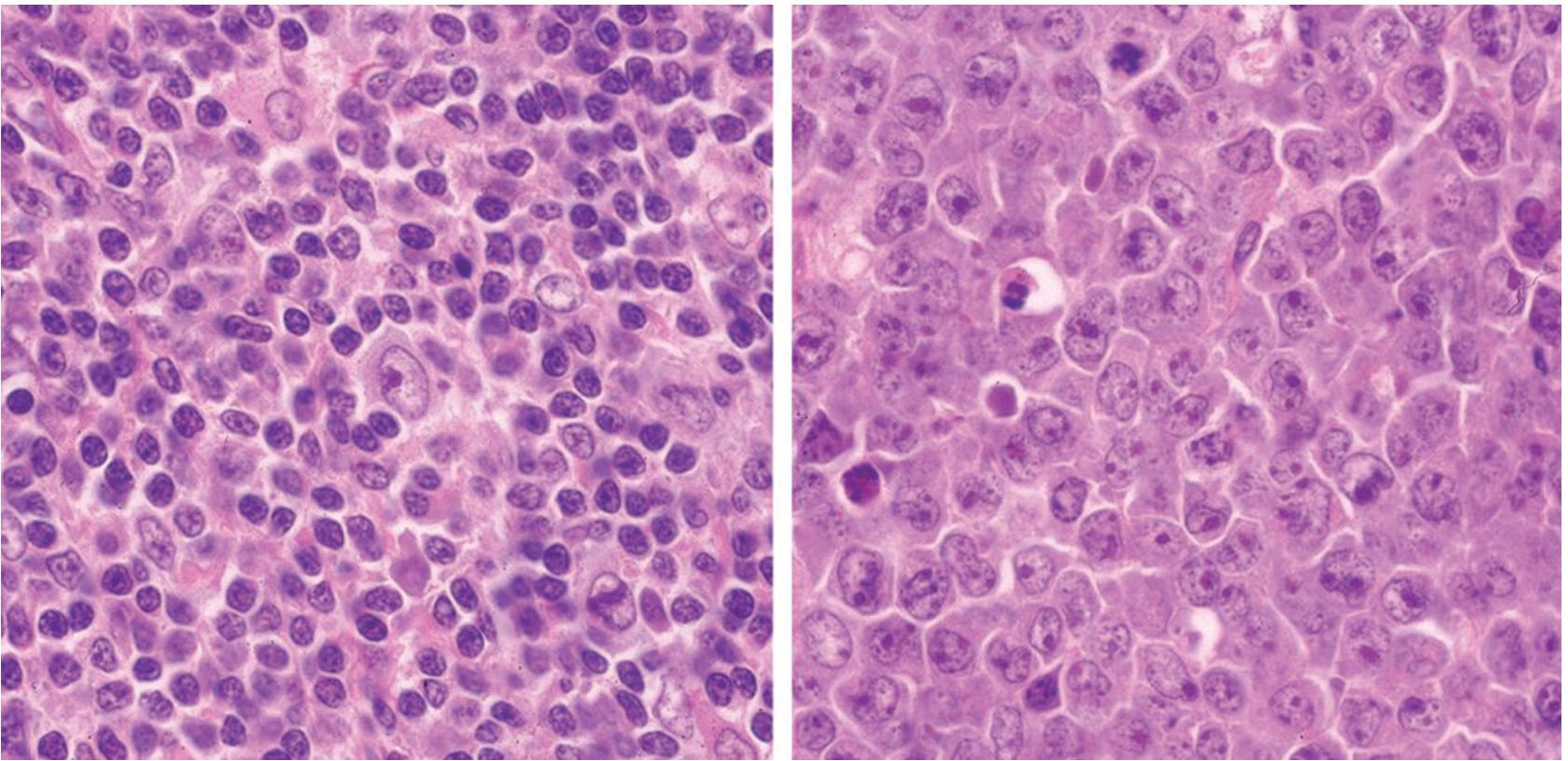
Figure 7 Composite lymphomas: angioimmunoblastic T-cell lymphoma (left) and non-Hodgkin lymphoma, diffuse large B-cell (right).
DISCUSSION
Although AITL (previously called angio-immunoblastic lymphadenopathy, AILD) was first described as a non-neoplastic lymphoid proliferation associated with immunodeficiency, it is clear that the authors of the Revised European-American Classification of Lymphoid Neoplasms4 regard this entity as a peripheral T-cell lymphoma because most cases show clonal rearrangements of TCR genes. In the past decade, there have been many reports of IgH clonal rearrangements in AITL, up to 40% of reported cases10,11. B cell and T cell clonal proliferations also have been reported in patients with PTCL-NOS, IgH clonal rearrangements were detected in 35% of PTCL-NOS patients20.
In this study we investigated cell lineages in 50 cases of AITL. TCR-γ monoclonal band/s were observed in 35 patients (70%), and IgH monoclonal band/s were observed in 16 patients (32%). Bilineage monoclonal proliferation of T and B cells was observed in 14 patients (28%). The presence of EBV genomes has no association with B cell clonality. We believe that ATTL is a dual malignancy which occurs simultaneously in a lymph node instead of being a T cell neoplasm only. This hypothesis is supported by various reports of DLBCL arising in patients with preexisting AITL13,21,22, and composite lymphomas of AITL and DLBCL12. The above reported cases were EBV-associated with the possibility that the EBV-associated DLBCL was the secondary event in the AITL via EBV infection or reactivation followed by clonal expansion of an immortalized EBV-infected B cell clone12. Other secondary lymphomas that occurred in AILT patients included small B-cell lymphomas, plasmacytoma, PTCL-NOS, and classical Hodgkin lymphoma2324. Our three additional patients with composite lymphomas confirmed the neoplastic nature of T and B cells in AITL which progressed to mature T-cell lymphoma or mature B-cell lymphoma, the transformation may or may not be associated with latent EBV infection. For the above reasons, AITL is considered to be a unique disease entity which should not be classified in either mature T-cell lymphoma or mature B-cell lymphoma.
The data presented here may indicate association of AITL with the latent EBV infection since EBERs positivity was detected in 46% of our cases which is somewhat lower than most reports from other countries (which vary from 37% to 97%)6,7,25,26, which may be partially explain by noting that we used strict criteria for the interpretation of EBERs positivity. A few positive cells or positive less than 5% of the total tumor cell population were considered to be negative. We chose this strict standard, as a previous study we did showed that a small amount (less than 5%) of EBERs-positive cells was detected in 46% of nodal lymphoid hyperplasia cases5, a finding concurred with a previous report in which EBV DNA in peripheral blood CD3- lymphocytes was detected in 50% of the healthy individuals27. These EBERs-positive cells were considered to be by-stander lymphocytes, and thus we decided to exclude them from our analysis, which may be resulted in the lower findings than in most other studies.
The clinical course in this group of patients was aggressive with a median survival of 36 months despite chemotherapy in most cases. Our median and overall survival times were better than in one reported European series14, which may have been due to younger age of our patients overall, with 10% being teenagers. The expression of EBERs in tumor cells had no association with the disease outcomes.
Overall, the two populations of classical pattern and large-cell rich pattern had no statistically significant difference in their survival profiles. A previous report also documented insignificant differences in a similar population in terms of clinical features, laboratory findings and treatment response14. The morphologic heterogeneity and lack of specific immunophenotypic marker for AITL could be a source of misdiagnosis to inexperienced pathologists, especially cases of AITL with an interfollicular pattern, AITL with very rich in epithelioid cells, and AITL with a granulomatous formations. The morphologic patterns have no clinical or prognostic values, but may be helpful in guiding pathologists to make the correct diagnosis of this disease.
ACKNOWLEDGMENTS AND REMARKS
This work was supported by a research grant from Faculty of Medicine, Prince of Songkla University. The study was approved by the Hospital Ethics Committee, Faculty of Medicine, Prince of Songkla University.
REFERENCES
1. Swerdlow SH, Compo E, Harris NL, Jaffe ES, Pileri SA, Stein H, Thiele J, Vardiman JW (Eds.): WHO classification of Tumours of Hematopoietic and Lymphoid Tissues. IARC: Lyon 2008:309-311.
2. Frizzera G, Moran EM, Rappaport H. Angioimmunoblastic lymphadenopathy with dyspro-teinaemia. Lancet 1974;1:1070-1073.
3. Dogan A. Angioimmunoblastic T-cell lymphoma: review. Brit J Haematol 2003;121:681-691.
4. Harris NL, Jaffe ES, Stein H, et al. A revised European-American classification of lymphoid neoplasms: a proposal from the International Lymphoma Study Group. Blood 1994;84:1361-1392.
5. Mitarnun W, Pradutkanchana J, Ishida T. Ep-stein-Barr virus-associated nodal malignant lymphoma in Thailand. Asian Pacific J Cancer Prev 2004;5:268-272.
6. Lee Y, Lee KW, Kim JH, et al. Epstien-Barr virus-positivity in tumor has no correlation with clinical outcomes of patients with angioinmuno-blastic T-cell lymphoma. Korean J Intern Med 2998;23:30-36.
7. Anagnostopoulos I, Hummel M, Tiemann M, et al. Heterogenous Epstein-Barr virus infection pattern in peripheral T-cell lymphoma of angioimmunoblastic lymphadenopathy type. Blood 1992;80:1804-1812.
8. Lorenzen J, Li G, Zhao-Hohm M, et al. Angioimmunoblastic lymphadenopathy type of T-cell lymphoma and angioimmunoblastic lymphad-enopathy: a clinicopathological and molecular biological study of 13 Chinese patients using polymerase chain reaction and paraffin-embedded tissues. Virchows Arch 1994;424:593-600.
9. Feller AC, Griesser H, Schillina CV, et al. Clonal gene rearrangement patterns correlate with immunophenotype and clinical parameters in patients with angioimmunoblastic lymphad-enopathy. Am J Pathol 1988;133;549-556.
10. Willenbrock K, Roers A, Seidl C, et al. Analysis of T-cell subpopulations in T-cell non-Hodgkin’s lymphoma of angioimmunoblastic lymphadenopathy with dysproteinemia type by single target gene amplification of T cell receptor-beta gene rearrangements. Am J Pathol 2001;58:1851-1857.
11. Smith JL, Hodges E, Quin CT, et al. Frequent T and B cells oligoclones in histologically and immunophenotypically characterized angioimmunoblastic lymphadenopathy. Am J Pathol 2000;156:66-69.
12. Xu Y, McKenna RW, Hoang MP, et al. Composite angioimmunoblastic T-cell lymphoma and diffuse large B-cell lymphoma: a case report and review of the literature. Am J Clin Pathol 2002;118:848-854.
13. Hawley RC, Cankovic M, Zarbo RJ. Angioim-munoblastic T-cell lymphoma with supervening Epstein-Barr virus-associated large B-cell lymphoma. Arch Pathol Lab Med 2006;130:1707-1711.
14. Mourad N, Mounier N, Briere J, et al. Clinical biologic and pathologic features in 157 patients with angioimmunoblastic T-cell lymphoma treated within the Group’e d’Etude des Lymphomes de I’Adulte (GELA) Trials. Blood 2008;111:4463-4470.
15. Attygalle A, Al-Jehani R, Diss TC, et al. Neoplastic T cells in angioimmunoblastic T-cell lymphoma express CD10. Blood 2002;99:627-633.
16. Ree HJ, Kadin ME, Kikuchi M, et al. Angioim-munoblastic lymphoma (AILD-type T-cell lymphoma) with hyperplastic germinal centers. Am J Surg Pathol 1998;22:643-655.
17. Kieff E, Leibowitz D. Epstein-Barr virus and its replication. In: Field BN, ed. Viology, 2nd edn. New York: Raven Press. 1990;1889-920.
18. McCarthy KP, Sloane JP, Kabarowski JHS, et al. A simplified method of detecting clonal rearrangements of the T-cell receptor-g chain gene. Diagn Mol Pathol 1992;1:173-179.
19. Amara K, Trimeche M, Sriha B, et al. PCR-based clonality analysis of B-cell lymphomas in paraffin-embedded tissues: diagnostic value of immunoglobulin kappa and lambda light chain gene rearrangement investigation. Pathol Res Pract 2006;202:425-431.
20. Tan BT, Warnke RA, Arber DA. The frequency of B- and T-cell gene rearragements and Ep-stein-Barr virus in T-cell lymphomas. A comparison between angioimmunoblastic T-cell lymphoma and peripheral T-cell lymphoma, unspecified with and without associated B-cell proliferations. J Mol Diagn 2006;8:466-475.
21. Abruzzo LV, Schmidt K, Weiss LM, et al. B-cell lymphoma after angioimmunoblastic lymphade-nopathy: a case with oligoclonal gene rearrangement associated with Epstein-Barr virus. Blood 1993;82:241-246.
22. Knecht H, Maritus F, Bachmann E, et al. A deletion mutant of the LMP1 oncogene of Epstein-Barr virus evolution of angioimmunoblastic lymphadenopathy into B immunoblastic lymphoma. Leukemia 1995;9:458-465.
23. Warnke RA, Jones D, Hsi ED. Morphologic and immunophenotypic variants of nodal T-cell lymphoma and T-cell lymphoma mimics. Am J Clin Pathol 2007;127:511-527.
24. Attygalle AD, Kyriakou C, Dupis J, et al. Histologic evolution of angioimmunoblas-tic T-cell lymphoma in consecutive biopsies: clinical correlation and insight into natural history and disease progression. Am J Surg Pathol 2007;31:1077-1088.
25. Weiss LM, Jaffe ES, Liu XF, et al. Detection and localization of Epstein-Barr viral genomes in angioimmunoblastic lymphadenopathy and angioimmunoblastic-like lymphoma. Blood 1992;79:1789-1795.
26. Noorali S, Pervez S, Moatter T, et al. Characterization of T-cell non-Hodgkin’s lymphoma and its association with Epstein-Barr virus in Pakistani patients. Leuk Lymphoma 2003;44:807-813.
27. Mitarnun W, Suwiwat S, Pradutkanchana J, et al. Epstein-Barr virus-associated peripheral T-cell and NK-cell proliferative disease/lymphoma: a clinicopathologic, serologic and molecular analysis. Am J Hematol 2002;70:31-38.


